3.2. Extraction efficiency for molecular methods
Sample collection volumes, types, and preservation methods varied throughout the collection time period as sampling and extraction protocols were still being optimized. As such, DNA quantity in extractions was highly variable due to the variety in the sample state, density, and preservation methods applied. Cultured samples gave the highest yields. Filtered field samples generally resulted in lower, but adequate yields. Pelleted field samples resulted in low and often unusable yields. After extractions, 46 of the 54 samples had sufficient high-quality DNA for molecular analyses. Samples without sufficient DNA (i.e., s025, s026, s027, s029, s039, s051, s052, s054) were not run on qPCR, but two of these samples (i.e., s029 and s054) were run through metabarcoding NGS and returned operational taxonomic units (OTUs) (<25 and <1350, respectively). Five samples (i.e., s034, s045, s046, s050, s053) had low DNA concentration, but returned positive results with qPCR and NGS.
3.3. Taxa identified by qPCR
A total of 46 samples with sufficient DNA concentrations were run using qPCR. However, only seven of 39 applied TaqMan assays provided amplification with one or more of the samples (
Table S2), and the rest (
n = 32 assays) did not return positive or suspected results in any samples. These seven assays amplified within 43 of the samples. Targeted species that were detected via qPCR included
Alexandrium spp./
A. tamarense,
H. akashiwo, and
P. verruculosa. The targeted
Chattonella spp. had four different qPCR assays, but none resulted in detections. Other species that are potentially harmful were identified by qPCR, but were not specifically identified by microscopy. For example, cyanobacteria species were detected in nine samples and
Karlodinium micrum/
veneficum in ten samples. Nine targeted taxa were not quantifiable by qPCR due to a lack of published TaqMan assays:
C. concavicornis,
C.
convolutus,
Chrysochromulina spp.,
Margalefidinium fulvescens,
Dictyocha spp.,
Dinophysis spp.,
P. reticulatum, and
Pseudo-nitzschia spp. Because this study ran the assays under different conditions than they were designed for, absence of positive controls, and because of algae strain variability, interpretation of qPCR results here was limited. NGS was conducted to confirm if the sample contained a target species, and therefore to determine the effectiveness of the tested assays and correspondence to microscopy. The relative effectiveness of the qPCR results here in comparison to microscopy and metabarcoding is provided in the Discussion (section 4).
3.4. Metabarcoding overview
In total, over 350 individual taxa were detected by metabarcoding NGS. NGS returned results for 48 of 48 sequenced samples. Detailed results of the analysis including all taxa and associated read counts are provided in
Tables S3–
S7.
The most numerous unique taxa occurred through using the 18S-dinoflagellate amplicon (
Table 2), which returned 167 individual taxa, including 49 to the species level. For comparison, the 16S amplicon identified 69 individual taxa (21 to the species level), and the 18S-diatom amplicon identified 78 taxa (18 to the species level). The 28S data identified 136 individual taxa and was the most effective at resolving data to the species level (
n = 60 species), which is noteworthy given the importance of species-level identification when identifying harmful algae. Only one taxon,
H. akashiwo, was identified by all four amplicons. Many similar taxa (
n = 20) were detected by 18S-diatom, 18S-dinoflagellate, and 28S amplicons, but only three were identified to the species level.
The number of reads per taxonomic category, with an emphasis on microalgal groups, are listed in
Table 3. Total microalgae reads detected by 18S-diatom, 18S-dinoflagellate, and 28S runs were more than 85% of all reads (not including unknowns) for each amplicon, whereas microalgae reads from 16S run comprised <50%. The majority of microalgal reads for all amplicons belonged to raphidophytes. The 18S-dinoflagellate amplicon detected the most microalgal groups and the 16S amplicon detected the fewest microalgal groups. Silicoflagellates (Dictyochophyceae), an important group that includes several harmful and potentially harmful species, were detected only by 18S-diatom and 18S-dinoflagellate amplicons.
All harmful and potentially harmful algal taxa reads (i.e., at the species, genus, and family level) detected by all amplicons are provided in
Table 4. For a more convenient comparison between results employing different amplicons, grouping of harmful and potentially harmful algae species to genus level was done for
Alexandrium spp.,
Chattonella spp.,
Dinophysis spp.,
Karlodinium spp.,
Phalacroma spp.,
Prymnesium spp.,
Pseudo-nitzschia spp., and
Pseudochattonella spp.
Overall, most of the reads of harmful and potentially harmful algae were detected by 28S and 18S-dinoflagellate amplicons (
Table 4). At the species and genus levels, the 28S amplicon provided the most reads for dinoflagellate species within the
Margalefidinium,
Dinophysis,
Karlodinium, and
Phalacroma genera, as well as haptophytes in the
Chrysochromulina and
Prymnesium genera. The 18S-dinoflagellate amplicon detected the most reads for species within the
Alexandrium,
Pseudochattonella, and
P. reticulatum genera, whereas the 18S-diatom amplicon detected the most reads for
Pseudo-nitzschia spp. as well as the raphidophytes
Chattonella sp. and
H. akashiwo. There were no instances when 16S detected the most reads for any of the listed taxa.
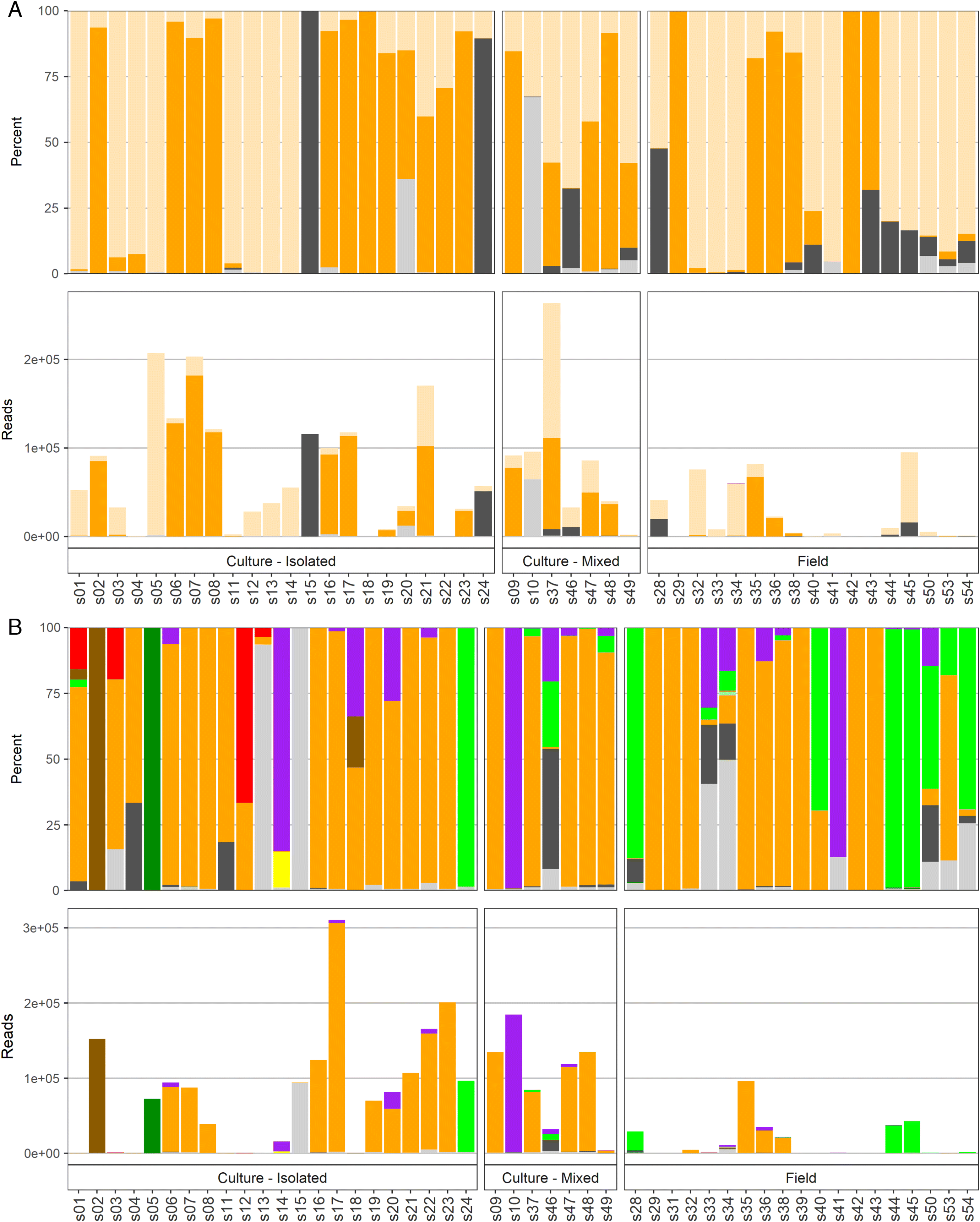
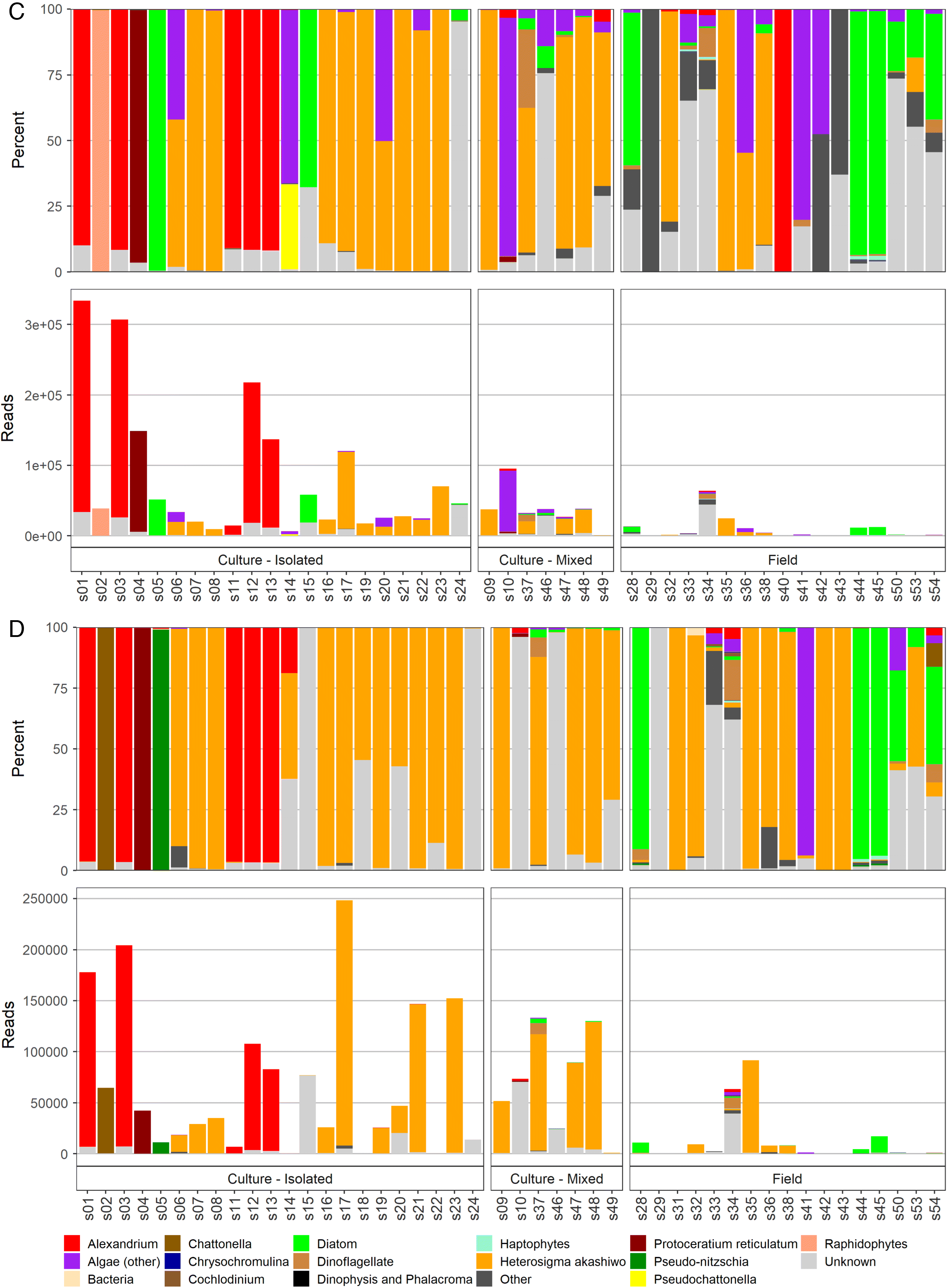
The total reads and percentages per sample detected by different amplicons are shown in
Fig. 2. Specifically in this plot, when a taxon of harmful or potentially harmful algae had <100 reads, it was grouped within an appropriate, larger algae category, e.g.,
Karlodinium spp. with 77 reads were included in the dinoflagellates counts;
Prymnesium spp. with 20 reads and Prymnesiaceae with 11 reads were included in haptophytes (
Fig. 2). A comparison of microscopy and NGS taxonomic identification in each of the 48 samples revealed that the majority of the taxa detected by microscopy also were identified in the NGS results (
Table S7). Species and genera that were positively identified by both microscopy and NGS included:
Alexandrium,
Chaetoceros,
Margalefidinium,
H. akashiwo,
P. reticulatum,
Pseudochattonella, and
Pseudo-nitzschia. There were a few notable exceptions, with the two most consistently observed mismatches being (
i) all suspected
Chattonella spp. (four field and six mixed cultures) were not confirmed as Chattonella and (
ii)
Dictyocha spp. (six field samples),
P. verruculosa (one culture),
Pseudochattonella sp. (five field samples), and
Pseudopedinella spp. (one culture and five filed) (i.e., all three silicoflagellate genera) were not detected by any amplicon in 11 out of 15 samples suspected to contain these taxa based on microscopy. Although these samples had unknown and unassigned reads in NGS results (
Table S7), the sequences for all of the species not identified by NGS but identified with microscopy were in fact present in the GenBank database; therefore, the reason for the lack of these taxa, as well as the identity of these reads remains unknown.