Biotic variables
Abiotic and biotic species trait data (i.e., life history characteristics, habitats and threats) were extracted from a variety of publicly accessible online databases for birds (
Table S1;
Myhrvold et al. 2015;
Sheard et al. 2020), fishes (
Table S2;
Froese and Pauly 2016), mammals (
Table S3;
Wilman et al. 2014;
Myhrvold et al. 2015), and amphibians and reptiles (
Table S4;
Myhrvold et al. 2015;
Oliveira et al. 2017;
Santini et al. 2018;
Grubler 2020). Data extraction was conducted separately for each taxon, given differences among ideal databases, common variables, and units reported across taxonomic groups. For instance, length was typically recorded for fishes and herpetofauna (
Froese and Pauly 2016,
Santini et al. 2018), while mass was more commonly reported for birds and mammals (
Myhrvold et al. 2015). For each taxonomic group, variables with sufficient data were mapped onto species from the C-LPI according to binomial scientific name using the
traitdata (
RS-eco 2021; birds, mammals, herpetofauna),
squamata (
Grubler 2020; herpetofauna),
and fishbase (
Boettiger et al. 2012; fishes) R packages—creating multi-variable datasets that are now publicly available for future analyses. The code and extracted data for Canadian vertebrates can be found online (
Currie et al. 2022). Note that only body size, lifespan, and trophic level were included in our analysis as they had broad taxonomic coverage, but additional traits with sufficient species coverage were also extracted for public interest and use. Moreover, the available code can be adapted to extract additional traits of interest from the available databases. Details on data extraction for biotic variables with sufficient species coverage, by taxon, are found below.
Birds
Five biotic variables were selected for inclusion within the C-LPI Bird Trait database (
Table S1), based upon data availability. Using the R package
traitdata (
RS-eco 2021), adult body mass, lifespan (maximum longevity), and mean longevity were extracted from the Amniote database (
Myhrvold et al. 2015). Data on body mass were also available from the EltonTraits database (
Wilman et al. 2014), but the Amniote database (
Myhrvold et al. 2015) was selected as the primary source given the quality of its metadata (i.e., more extensive). Average values per species were calculated for each biotic variable. In addition, hand-wing index (i.e., a measure of wing aspect ratio and a proxy for dispersal ability) and dietary guild were extracted from the Global Hand-Wing Index repository (
Sheard et al. 2020). Dietary guild categories were aligned to trophic level categorization (carnivore, omnivore, and herbivore). One species (turkey vulture,
Cathartes aura) was assigned to the carnivore category as it almost exclusively feeds on carrion.
Fishes
Data were extracted from FishBase (
Froese and Pauly 2016) using the R package
rfishbase (
Boettiger et al. 2012). Fourteen biotic variables were selected for inclusion within the C-LPI Fishes Trait database (
Table S2), including length, weight, and lifespan. Average values per species were calculated for each biotic variable.
Mammals
Gestation period and lifespan (maximum longevity) were retrieved from the Amniote database (
Myhrvold et al. 2015) using the R package
traitdata (
RS-eco 2021), while body size and diet were extracted from EltonTraits (
Wilman et al. 2014). In total, four biotic variables were selected for inclusion within the C-LPI Mammal Trait database, including body mass, trophic level, gestation period, and lifespan (
Table S3). Mammal trophic level was calculated using diet composition, where herbivore was defined as species associated with an entirely vegetarian diet (e.g., plants, seeds, nectar, and fruit), carnivores were classified as species that consume other animals, and omnivores included species with mixed diets.
iv. Amphibians & Reptiles
Seven biotic variables were selected for inclusion within the C-LPI Amphibian and Reptile Trait database (
Table S4), including variables related to body size, diet, and reproduction. Data were extracted from AmphiBIO (
Oliveira et al. 2017), the Amniote database (
Myhrvold et al. 2015,
Meiri 2018;
Santini et al. 2018,
Atwood et al. 2020), and SquamataBase (
Grubler 2020) using
traitdata (
RS-eco 2021) and
squamatabase (
Grubler 2020) R packages. Trait data that were comparable across datasets (e.g., body mass) were hierarchically extracted. For example, body mass was first extracted from AmphiBIO (
Oliveira et al. 2017), then from the amniote life history database (
Myhrvold et al. 2015) for species that lacked data in AmphiBIO. Because multiple entries were available per species within the amphibian allometry database (
Santini et al. 2018), values with the highest sample sizes were selected for inclusion. Amphibians and reptiles were grouped together as herpetofauna to improve sample size for analysis.
We also endeavored to extract of abiotic variables for this analysis—including species’ geographic ranges and average topographic and climate variables within those ranges. However, due to a lack of data coverage, we did not include these variables within our analysis. Nevertheless, the code and associated data can be found online.
Analyzing the representation of biotic variables
Three biotic variables with sufficient cross-taxa information were selected from the taxon-specific datasets to create a more fulsome cross-taxa database. The selection of traits was dependent upon a combination of (
i) data availability within and across taxa and (
ii) considerations on the available evidence of relationships between a trait and population declines and (or) extinction risk. For instance, call frequency and positioning are traits used to evaluate amphibian tolerance to anthropogenic pressures (e.g.,
Liu et al. 2021) but are not applicable biotic traits for fishes.
Furthermore, biodiversity loss is not random (
Dirzo et al. 2014) and functional traits, specifically, covary with patterns in biodiversity trends (
Munstermann et al. 2021;
Dirzo et al. 2014;
Lee and Jetz 2010). Trait diversity can therefore result in a biased impact on ecosystem functioning (
Diaz et al. 2006). For instance, body size is often a predictor of species loss, with large-bodied species particularly vulnerable (e.g.,
Dirzo et al. 2014;
Seguin et al. 2014;
Cardillo et al. 2005;
Solan et al. 2004;
Bennett and Owens 1997), though there has been mixed evidence dependent on taxonomic group (e.g.,
Chichorro et al. 2019;
Kopf et al. 2016).
Etard et al. (2020) found that while traits for mammals and birds are generally well-studied, there are gaps in information associated with reptiles and amphibians, making cross-taxa analyses more difficult. Moreover, a recent meta-analysis evaluating species traits and extinction risk identified body size as the most studied biotic trait, followed by fecundity and diet (
Chichorro et al. 2019). Yet, while fecundity was often investigated, it was infrequently linked to patterns in biodiversity trends. Alternatively, longevity was less frequently studied, but had the second-highest biotic variable significance (following body size). Lifespan was also found to be an important predictor of extinction risk for freshwater megafauna (
He et al. 2020). Likewise, we selected body size, longevity, and trophic level (i.e., diet) for our analysis based on the availability of information within and across taxonomic groups, as well as the evidence pertaining to relationship among these traits and biodiversity trends. While continuous traits are preferred for evaluating functional diversity (
Laureto et al. 2015), the categorical trait of trophic level was applied to assess trends across taxonomic groups, based on the taxon-specific databases available.
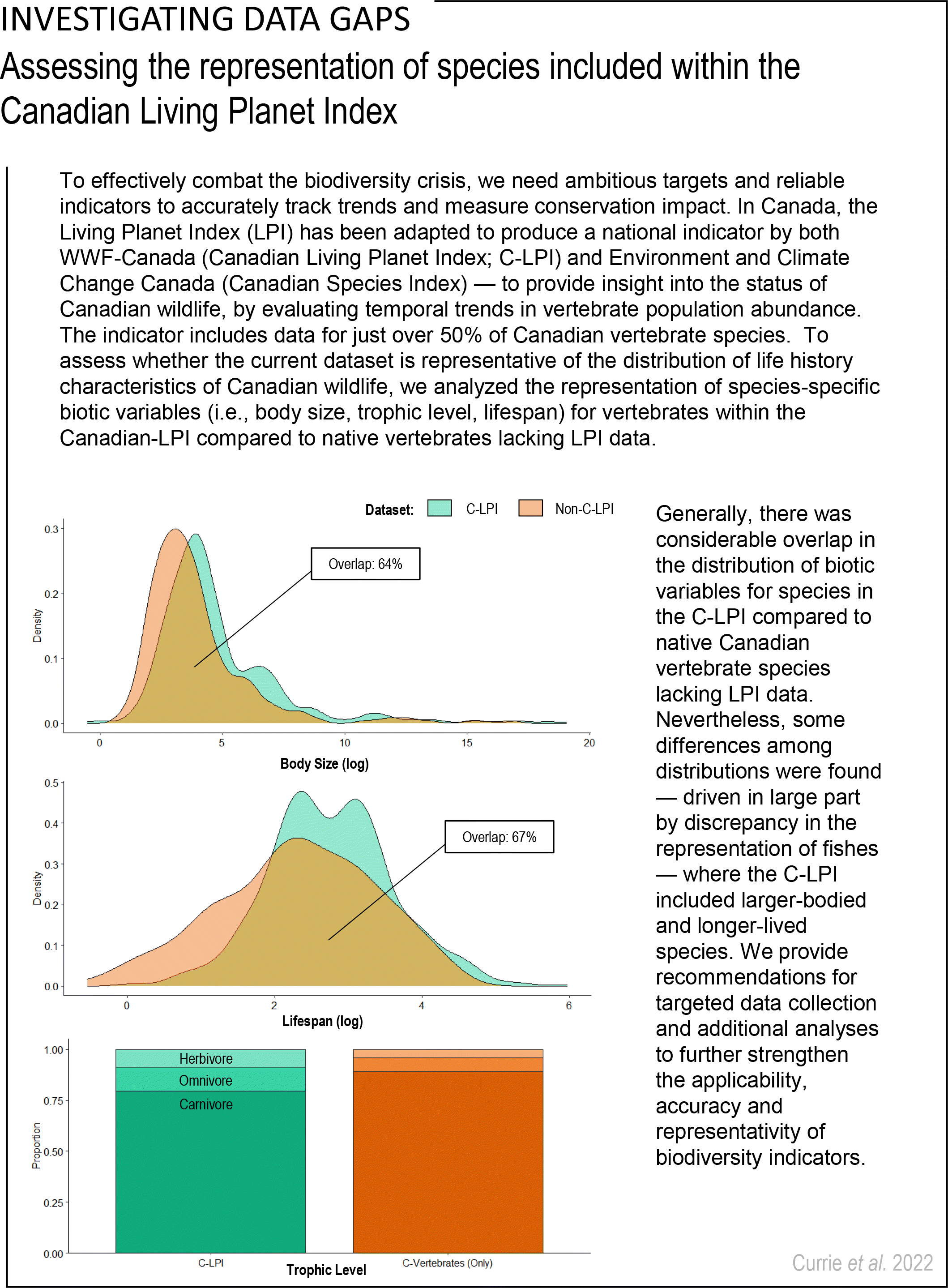
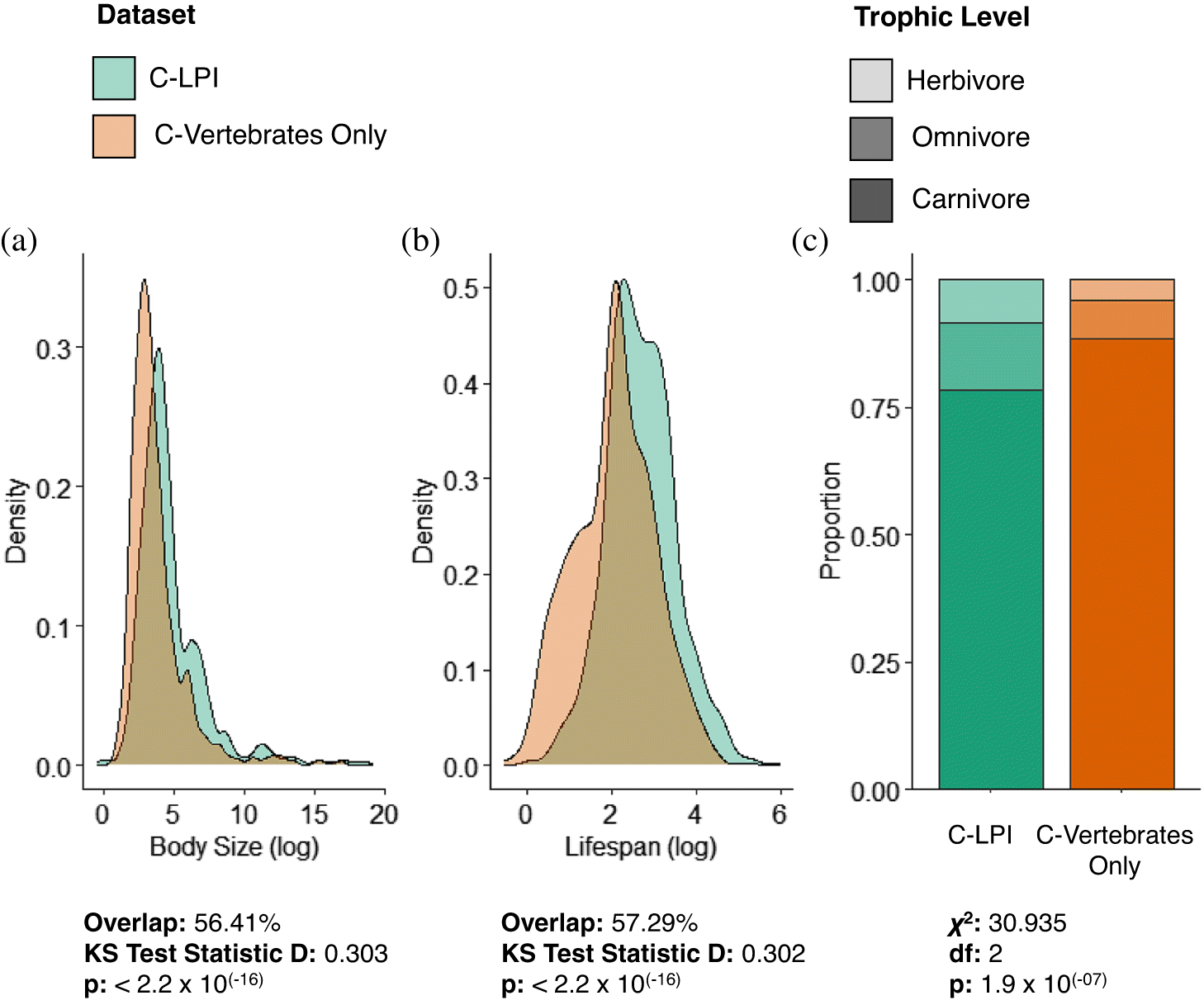
Continuous variables of body size and lifespan were log-transformed, while trophic level included three categorical levels (herbivore, omnivore, carnivore). The overarching cross-taxa C-LPI Trait Database included 836 species, while the broader C-Vertebrate Trait Database included 1679 species.
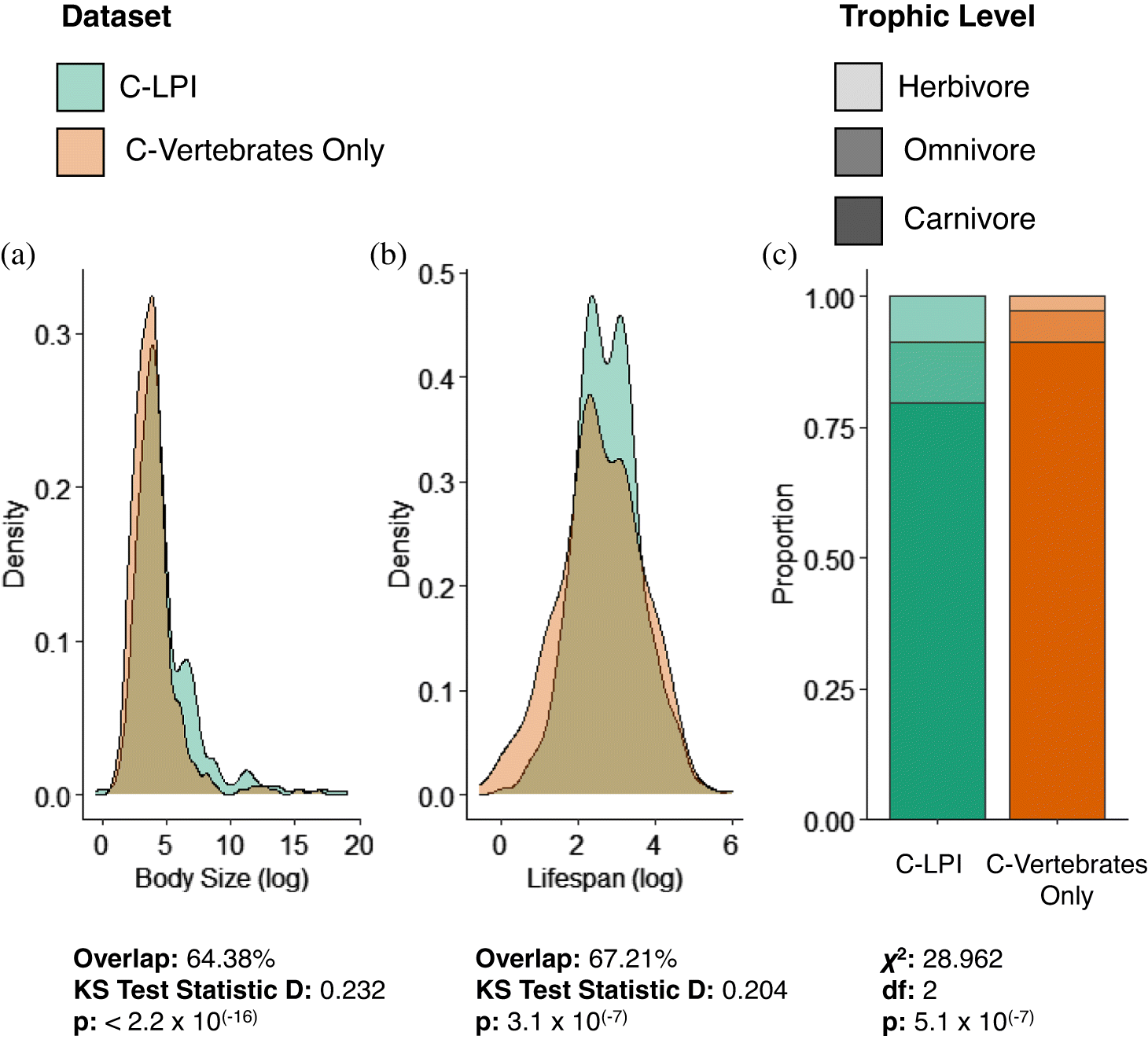
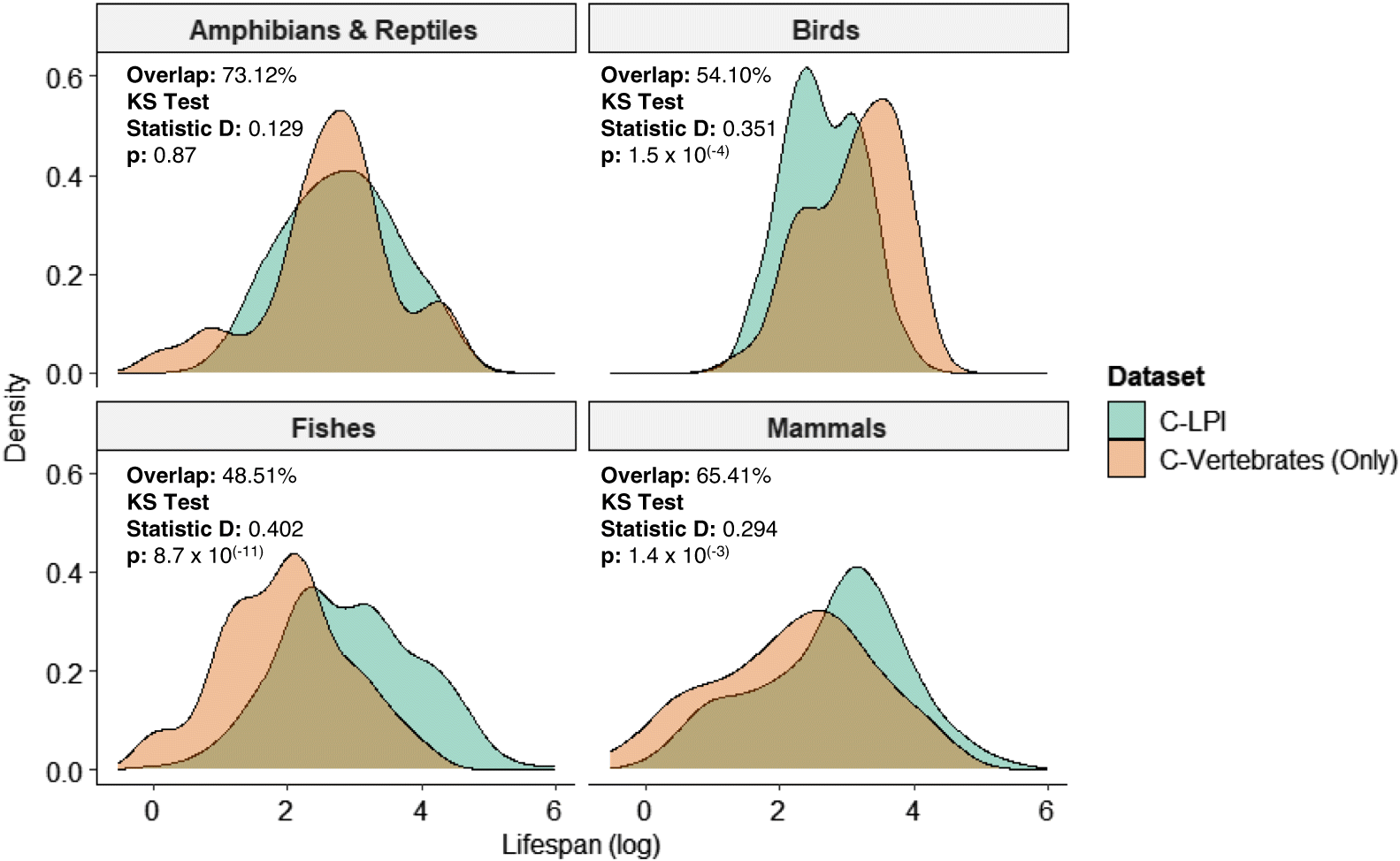
Shapiro–Wilk tests were used to assess assumptions of normality for the continuous variables of body size and lifespan for species included within the C-LPI trait database and for other native Canadian vertebrate species lacking LPI data (C-Vertebrates Only) (
Fig. S1). Given that the data generally exhibited skewed distributions, we applied two-sample nonparametric Kolmogorov–Smirnov (KS) tests to evaluate differences between the distribution of traits for species in the C-LPI Trait database and the C-Vertebrate (Only) Trait database. The widely used, asymptotic KS test measures distributional differences between two samples and has previously been applied to compare relationships among biotic variables (e.g.,
Adjeroud et al. 2007;
Langlois et al. 2012;
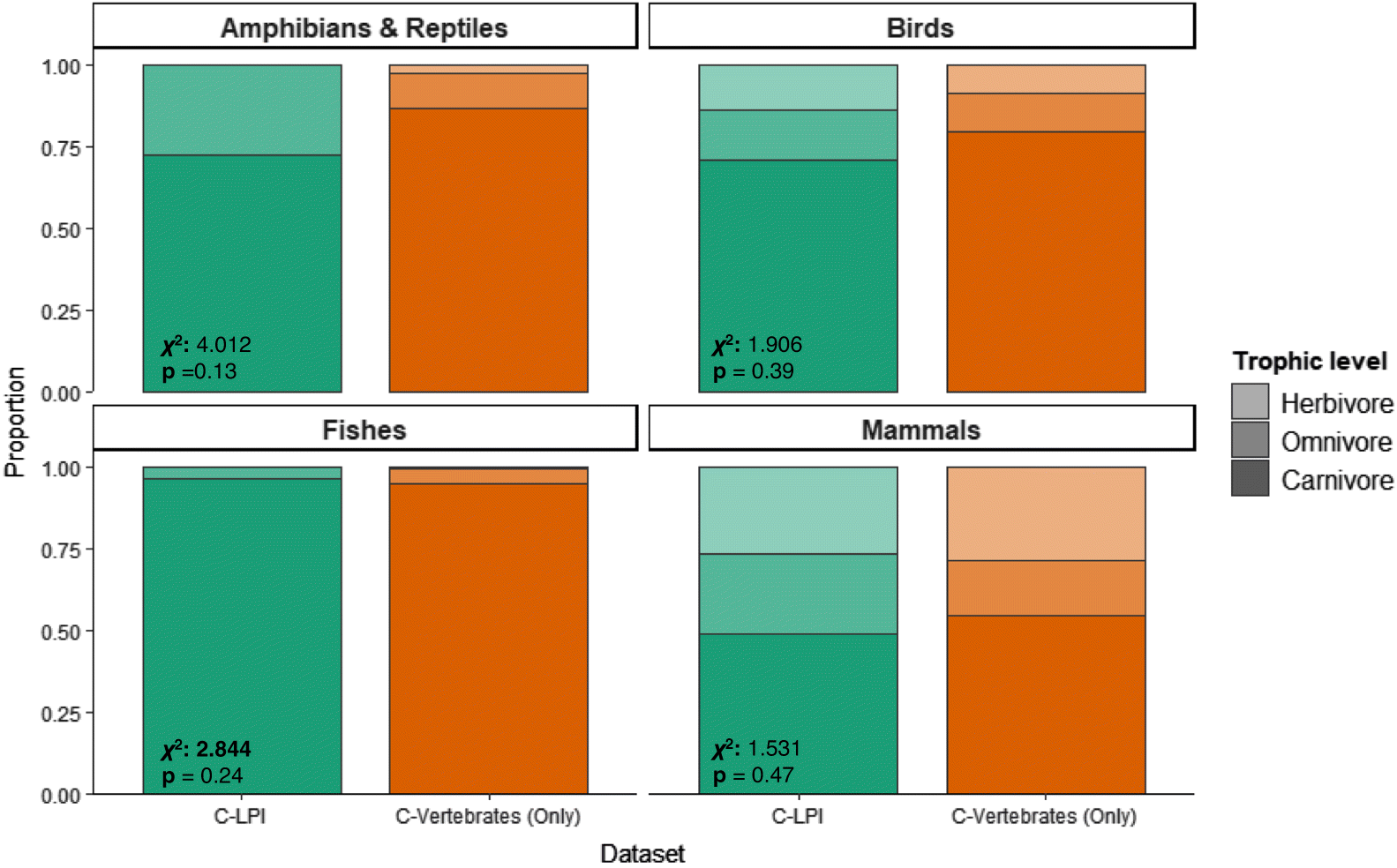
Cornwell et al. 2014). Importantly, we also assessed the percent overlap among distributions for continuous variables to evaluate the degree of differences in the distributions. These statistical tests were accompanied by a resampling approach to estimate differences in trait distributions between species in the C-LPI and C-Vertebrates (Only) datasets (see more below). We assessed the difference in categorical distribution of trophic level using Pearson’s χ
2 tests. Fisher’s exact test was used to validate small-sized samples that ran into computational errors during the application of Pearson’s χ
2 test (
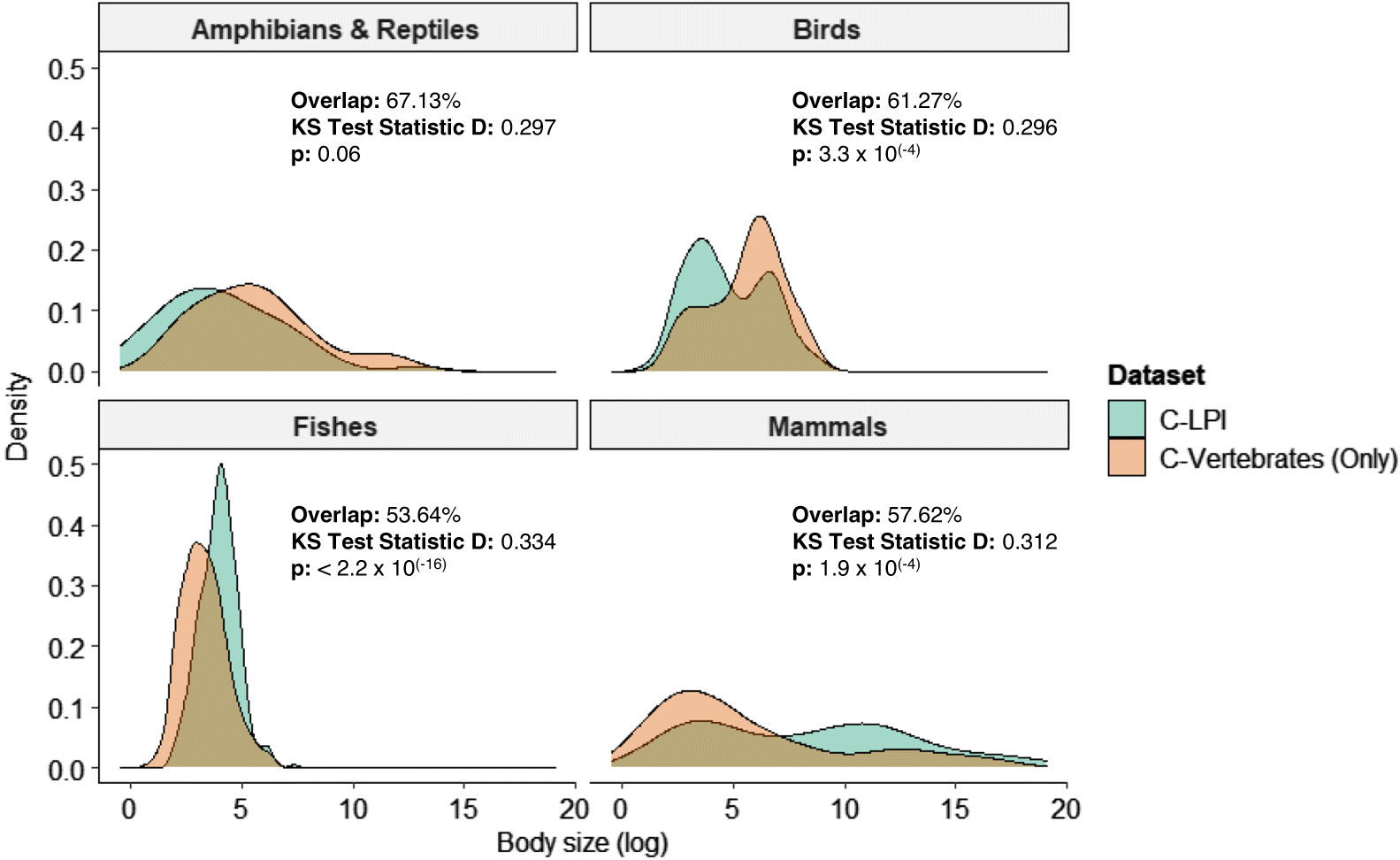
Kim 2017). All tests were performed for each biotic variable at the taxonomic level (i.e., birds, mammals, fish, amphibians and reptiles), and with all species combined.
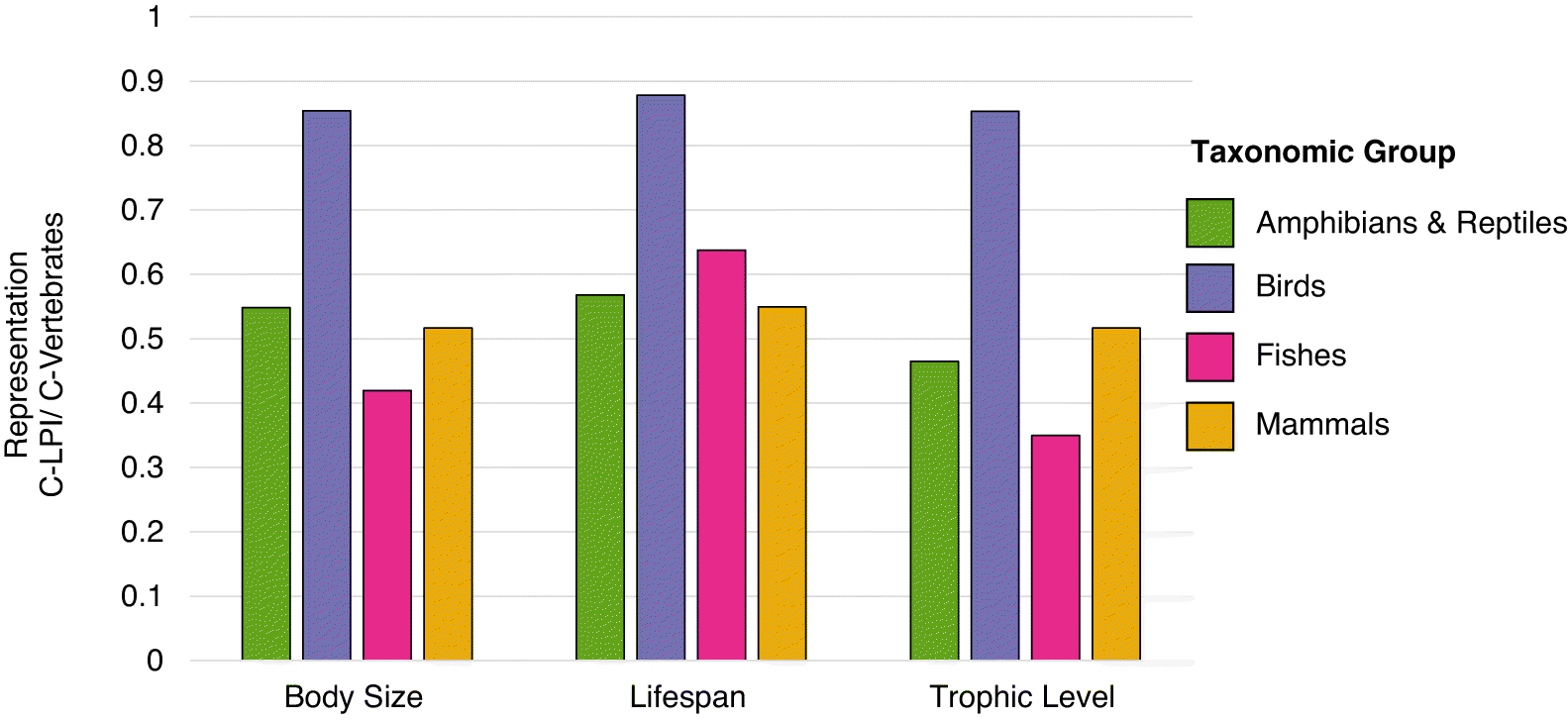
Completeness of variables in our dataset ranged between 35.0% and 87.8%. Consequently, we also reran the analysis with a more complete dataset, imputing missing biotic trait values via nonparametric imputation using the Random Forest approach (
Stekhoven and Buehlmann 2012) and an algorithm known as missForest (
Stekhoven 2013). This approach helped to identify important predictors of body size, lifespan, and trophic level using the additional biotic trait information that was extracted for each taxonomic group. The Random Forest approach permits biotic trait data imputation (i.e., estimation of missing data with complex relationships among multiple variables; e.g.,
Debastiani et al. 2021) and can deal with both continuous and categorical data to develop a more complete dataset for analysis (
Stekhoven and Buehlmann 2012).
To supplement our analysis of the two continuous traits (body size and lifespan), we applied a nonparametric bootstrapping approach to generate 95% compatibility intervals (95% CIs) on the median difference between datasets (i.e., the effect size;
Cohen 1994;
Ho et al. 2019). Bootstrapping was done via randomized resampling with replacement (
n = 5000 iterations) of the C-LPI and C-Vertebrate (Only) subsets of species-specific trait values to generate resamples equal in size to the original subsets. In addition, bootstrapping was performed twice: once using an unstratified approach in which all species were treated as a single sample without consideration for taxonomic representation and once using taxon-specific resampling in which data were sampled proportionally for each of the four vertebrate groups under consideration. We include the median and corresponding 95% CI on the difference between subsets for each of trait comparisons as a data-centered complement, and present Gardner–Altman estimation plots of these effects in the Supplementary Material.
All compiled trait data and associated code for this analysis are available through a Figshare repository (
Currie et al. 2022).